
With the continuous advancement of molecular oncology and precision medicine, homologous recombination deficiency (HRD) has emerged as a critical biological feature across several major solid tumors. Beyond ovarian cancer, a significant proportion of patients with breast cancer, prostate cancer and pancreatic cancer also present with HRD, BRCA1/2 mutations, or alterations in other homologous recombination repair (HRR) pathway genes. These defects impair the tumor’s ability to repair DNA double-strand breaks, rendering HRD-positive cancer patients more sensitive to PARP inhibitors and platinum-based therapies.
Clinical evidence indicates that:
・Around 50% of ovarian cancer cases show HRD positive;
・In breast cancer, germline BRCA1/2 mutations and broader HRR pathway alterations strongly predict PARP inhibitor sensitivity;
・In prostate cancer, BRCA1/2, ATM, PALB2 and other HRR gene mutations form the basis for regulatory approvals of PARP inhibitor therapies.
International guidelines now recommend HRD/HRR BRCA testing for ovarian, breast, pancreatic and prostate cancers to guide PARP inhibitor use, evaluate treatment response, and inform maintenance therapy strategies.
In response to this clinical demand, the Genecast PARPguide integrates the assessment of genomic instability (LOH, TAI, LST), BRCA1/2 variants, and additional HRR pathway gene alterations. This comprehensive approach supports precise patient stratification across multiple tumor types, enabling clinicians to more accurately identify individuals who are likely to benefit from PARP inhibitor therapy and tailor treatment decisions accordingly.
Genecast PARPguide integrates all critical biomarkers for identifying HRD status and guiding PARP inhibitor use:
・HRD Score: Based on LOH, TAI, and LST genomic scars
・BRCA1/2 Mutations: Germline and somatic, including large rearrangements
・Other HRR Pathway Genes: 64 genes panel for broader patient stratification
・Dual-platform strategy: Integrates low pass WGS and a SNP panel to enhance result stability.
・Low tumor requirement: Detects with as little as 20% tumor content, making testing accessible to more patients.
・High algorithm accuracy: Achieves 96% consistency between HRD score and BRCAness phenotype.
| Enrichment Method | Hybrid capture | |
| Panel Design | HRD score + 66 HRR (include BRCA1/2) | |
| Sequencing Platforms | NovaSeq 6000/NextSeq 500/550/DNBSEQ-T7/DNBSEQ-G400/DNBSEQ-G99 | |
| Package | 24 tests/kit | |
| Code | ZYCE-00701 | |

Patients newly diagnosed with ovarian, breast, pancreatic, or prostate cancer, particularly those being considered for platinum-based chemotherapy or PARP inhibitor therapy.

Individuals with recurrent or platinum-sensitive ovarian cancer undergoing evaluation for eligibility for maintenance therapy.

Individuals with a strong family history or clinical suspicion of hereditary cancer syndromes, to identify inherited risk or germline mutations.

Patients whose treatment strategy may benefit from HRD scoring or identification of HRR pathway gene mutations, including alterations in BRCA1/2, ATM, RAD51, PALB2, and related genes.
This test provides comprehensive molecular insights to assist clinicians in stratifying patients and optimizing personalized therapeutic strategies related to PARP inhibitors and DNA damage repair–targeted treatments.
| Sample Type | Sample Volume | Transport Conditions | Notes | |
| Whole Blood (EDTA tube) | Whole blood (≥8ml/tube). Insufficient volume may cause hemolysis due to negative pressure, which can affect testing accuracy. lt is recommended to send 2 tubes. | 6-37℃ | Postoperative blood must not be refrigerated or come into contact with ice packs. | |
| FFPE Tissue | FFPE Slides | Surgical tissue: ≥12 unstained slides (5-10 μm thick), total tissue area≥1 cm2 (approx, fingernail size). Biopsy: ≥15 unstained slides (5-10 μm thick), tissue area≥0.1 cm2 (approx, sesame size). | Room temperature or low temp, 0-30°C | Tumor content must be >20%. Rolls cannot be used for tumor cell proportion evaluation;please provide a set of slides for assessment. |
| FFPE Block | ||||
| FFPE Roll | ||||
| Surgical Tissue | Not less than 0.5cm x 0.5cm x 0.3cm, approx, the size of a soybean | Fresh tissue requires immersion in 10% neutral buffered formalin, and should reach the lab within 48 hours after resection. | ||
| Biopsy Tissue | ≥1cm in length, ≥3 pieces | |||
HRD scoring can be performed using different analytical platforms such as SNP panels, WES, or WGS, each with its own strengths and limitations. Genecast adopts an innovative dual-platform strategy combining Low-pass WGS and SNP panel analysis, achieving both uniform genomic coverage and high-density informative locus selection.
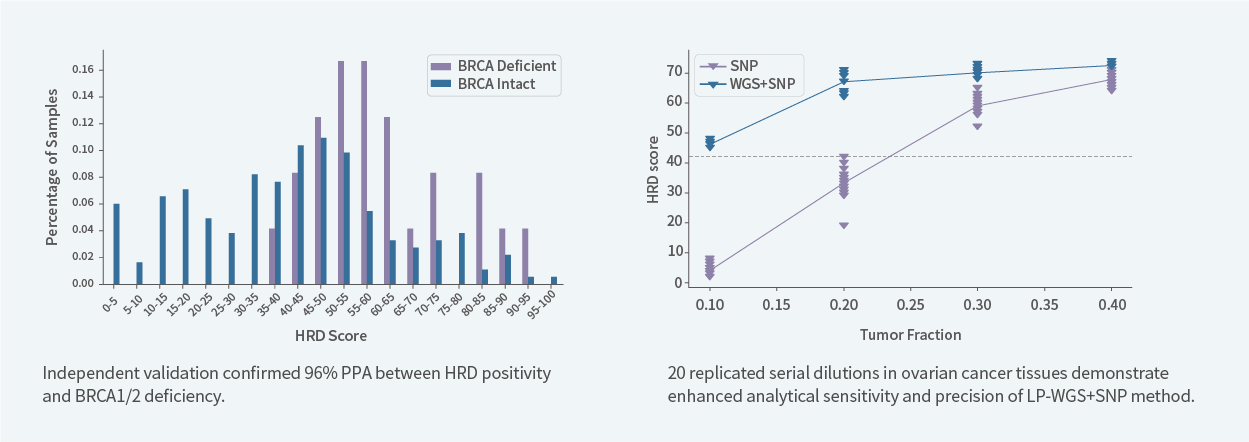
Validation using serially diluted ovarian cancer samples confirmed that the Low-pass WGS + SNP panel combined approach delivers higher sensitivity and precision for HRD scoring compared with SNP panel–only detection, and can reliably detect HRD when the tumor cellularity is as low as 20%.
Genecast PARPguide enables simultaneous detection of all three major biomarker categories associated with HRD including:
・HRD score (based on LOH, TAI, and LST)
・germline and somatic BRCA1/2 mutations (included LGR)
・Mutations in other HRR pathway genes
The patented HRD algorithm accounts for both BRCA-deficient and BRCA-intact tumors when determining classification thresholds. Consistency between HRD score–positive status and BRCA1/2 deficiency reaches 96%, demonstrating strong analytical reliability. Genecast’s proprietary genomic instability algorithm has been granted both domestic and international patents, ensuring robust and validated performance.

Lowpass WGS + SNP Panel Integration

Complete BRCA1/2 Reporting(inclue LGR)

5–7 Working Days Turnaround Time

Germline + Somatic Analysis