
Researchers have found that the characteristics of tumors are determined by their genetic makeup. Therefore, cancer treatment can begin at the genetic level by targeting specific genes to influence tumor behavior and achieve therapeutic outcomes. Tumor driver genes are associated with cancer progression and can help assess prognosis, as well as the risks of recurrence and metastasis. Drug-related genes provide valuable guidance for therapy selection by indicating drug sensitivity or resistance. Understanding the genetic profile of a tumor forms the basis for treatment planning. By identifying the appropriate genes, more precise and effective cancer therapies can be delivered, maximizing clinical benefit.
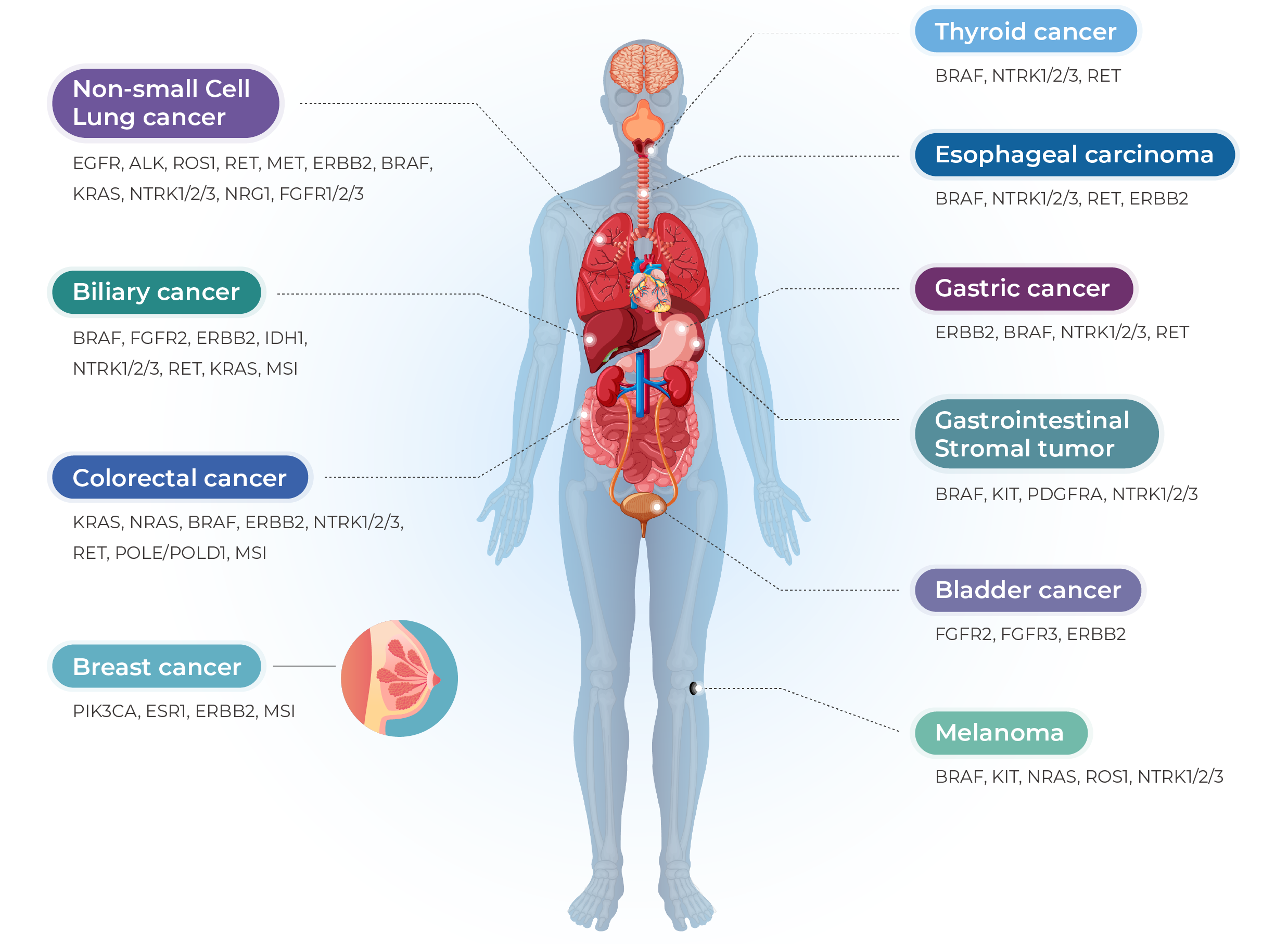
Genecast Comprehensive is a hybrid capture based test that uses targeted NGS to detect single nucleotide variants (SNVs), copy number variants (CNVs), insertions and deletions (InDels), and also reports a tumor mutational burden (TMB) score and microsatellite stability (MS) status in 769 genes using DNA extracted from FFPE tumor tissue samples from solid tumor patients. The test can also be used to detect fusions in 52 genes from RNA. Almost covering all solid cancer drivern genes and chemotherapeutic drug treatment related genes in NCCN guidelines. In addition, the test is intended to provide tumor profiling information to qualified healthcare professionals for solid tumor patients.
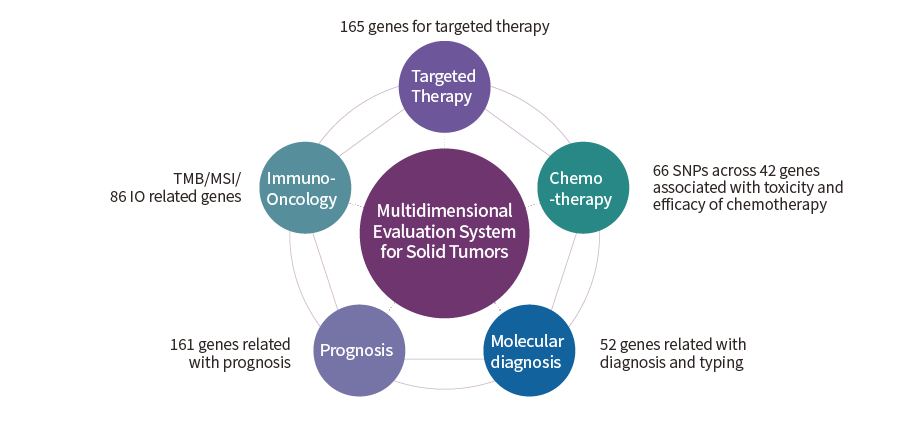
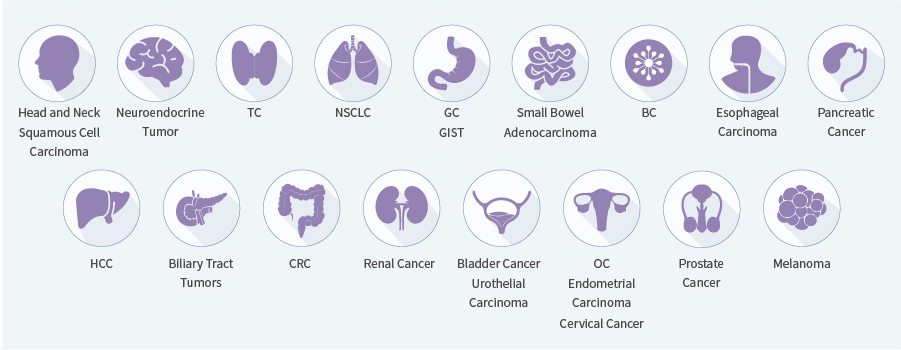
| Enrichment Method | Hybrid capture | |
| Panel Design | 785 genes + MSI, DNA 769 genes + RNA 52 genes | |
| Sequencing Platforms | DNBSEQ-G400/NextSeq 550 | |
| Package | 24 tests/kit | |
| Catalog Number | ZYCE-00201 | |
 Supports (Neo)Adjuvant and Systemic Therapy Decisions
Supports (Neo)Adjuvant and Systemic Therapy Decisions
Detects SNVs, CNVs, and InDels across 769 genes at the DNA level, and 52 clinically relevant fusions at the RNA level, enabling personalized treatment planning for both adjuvant and systemic settings.
 Guides Immunotherapy Suitability Assessment
Guides Immunotherapy Suitability Assessment
Reports TMB, MSI, and POLE/POLD1 mutation status to evaluate the likelihood of benefit from immune checkpoint inhibitors.
 Uncovers Resistance Mechanisms to Guide Subsequent Therapy
Uncovers Resistance Mechanisms to Guide Subsequent Therapy
Identifies key resistance mutations (e.g., EGFR T790M, KRAS G12C, HER2 amplification), providing molecular insights for optimizing second- and third-line treatment strategies.
 Assesses Recurrence Risk and Potential Residual Disease
Assesses Recurrence Risk and Potential Residual Disease
Detects high-risk mutations such as TP53, PIK3CA, and RB1 that may be associated with minimal residual disease or postoperative recurrence, supporting long-term follow-up planning.
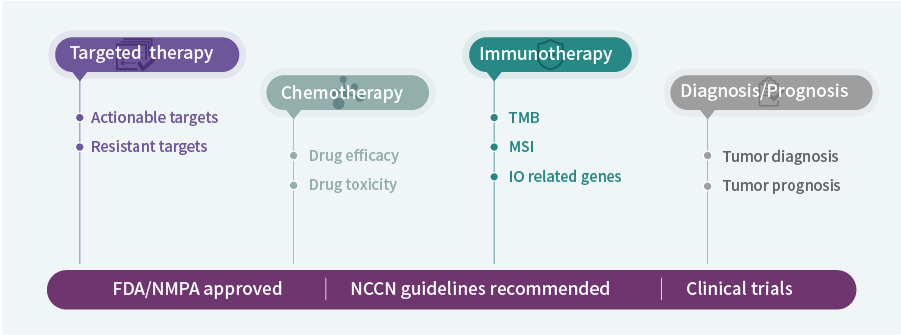
 Facilitates Clinical Trial Matching
Facilitates Clinical Trial Matching
Covers multiple NCCN/CSCO-endorsed biomarkers to assist in matching patients to targeted therapy or immunotherapy trials.


| Type | Requirements | Sample Quantity |
| Surgical/Large Biopsy Tissue Sections | Area ≥ 0.5 cm2, Thickness 5-10 μm; Tumor cell content ≥ 20%; Storage period ≤ 3 years. | ≥ 10 unstained slides/Wax rolls with 4 slices |
| Small Biopsy Tissue Sections | Area ≤ 0.5 cm2, Thickness 5-10 μm; Tumor cell content ≥20%; Storage period ≤ 3 years. | ≥ 12 unstained slides/Wax rolls with 6 slices |

DNA+RNA dual-assay

Detection of SNV / InDel/ CNV / Fusion / MSI / TMB

In-time update of Clinical Knowledge Base

Compatible with MGI and Illumina sequencing platforms